Which individual human factors favour the formation of biofilms?
What is this research project about?
Implant-associated infections represent a serious health burden in the clinics. They are the consequence of microorganisms that are able to colonise biological surfaces or surfaces of indwelling medical devices, and to form biofilms there. Biofilms are communities of microorganisms attached to such hydrated interfaces and enclosed in self-produced extracellular matrices. These structures present a considerable therapeutic challenge. Bacteria in biofilms exhibit altered phenotypes in their growth rates, gene expression profiles, and protein production. Embedded in their extracellular matrix, they become resilient to both antibiotics and the effector mechanisms of the immune system of the host. Since antibiotics treatment alone is apparently ineffective to treat these infections, the only possible treatment is the removal of the infected device followed by antibiotics. All of which results in high costs for both the patients and the healthcare facilities.
What’s the current status?
The connections between the formation of bacterial biofilms on implants and the immune system of the host have not been investigated in detail. Thus, still little is known on the role of the immune system in the development of implant-associated biofilms produced by S. aureus or P. aeruginosa or other pathogenic bacteria. However, for rational design of treatment strategies, this would be an essential asset.
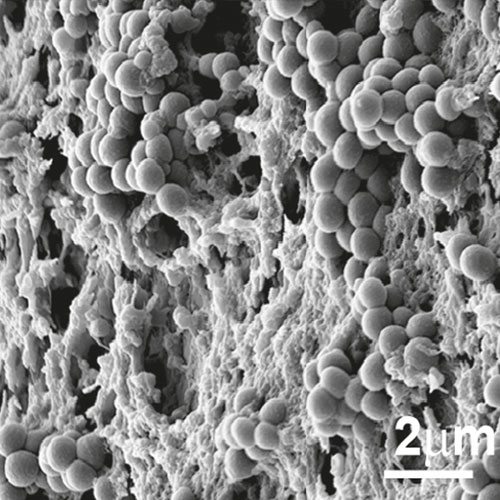
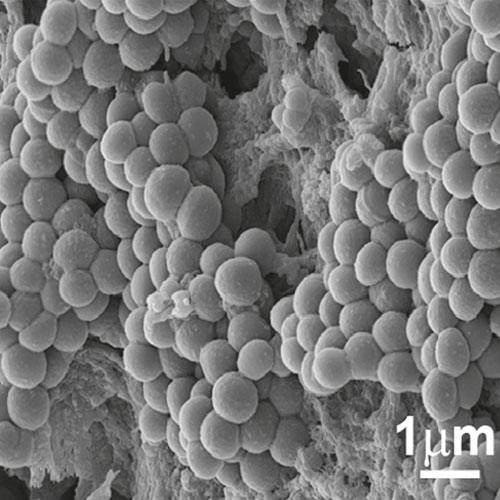
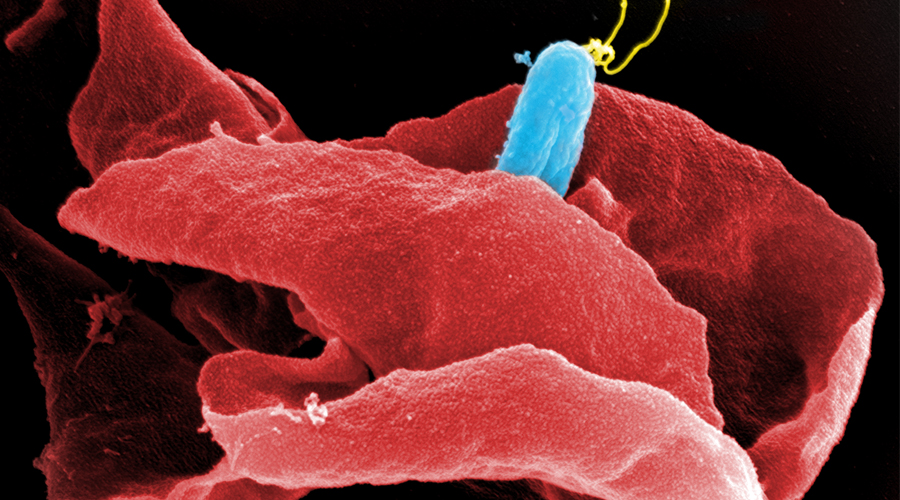
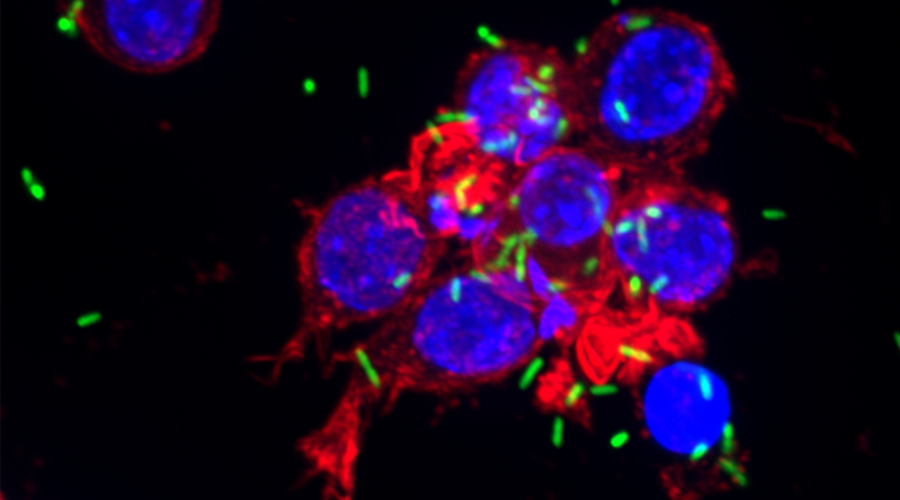
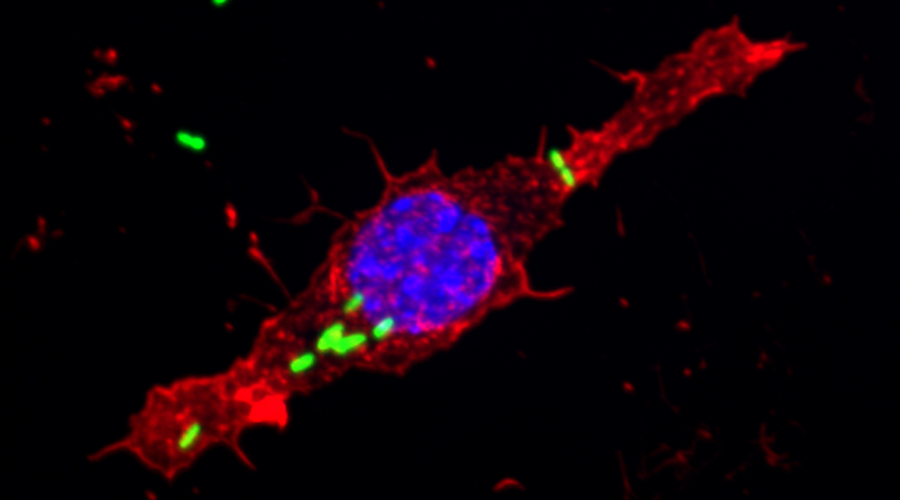
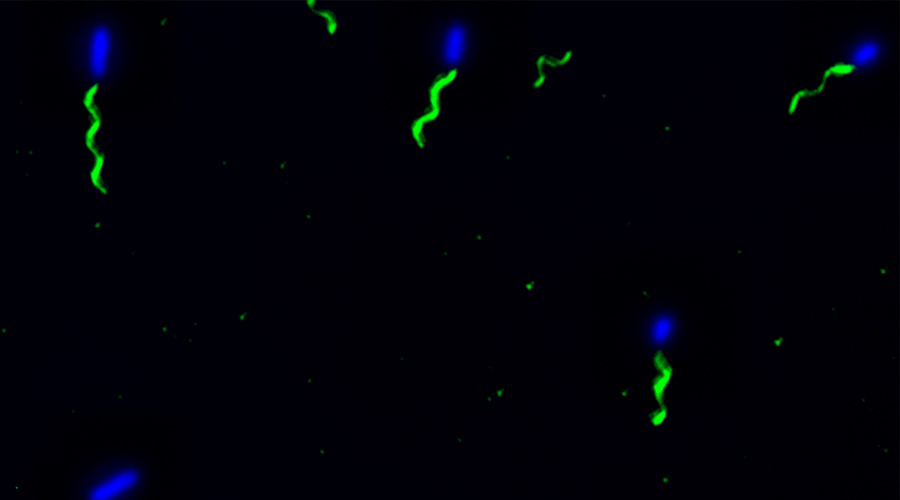
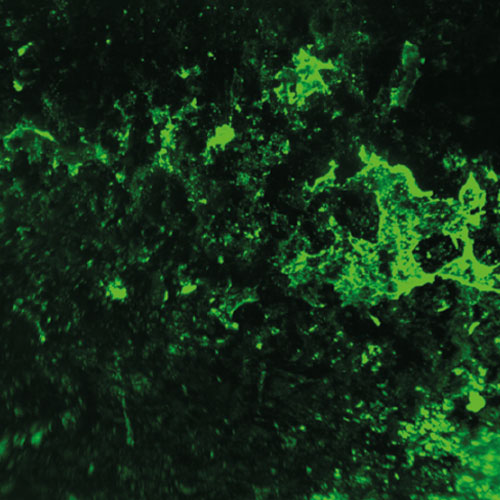
To develop a mouse model for studying implant-associated infections, we pre-colonized osmotic pumps with S. aureus and implanted them subcutaneously into C57Bl/6 mice. The bacteria expressed a bacterial luciferase which allowed monitoring of their expansion and the development of biofilms by employing noninvasive in vivo imaging. Subsequently, the presence of bacterial aggregates and biofilms on the surface of the implants was showed by immunofluorescence and electron microscopy. Additionally, myeloid cells, including neutrophils and monocytes, had been attracted to the colonized pumps and formed a barrier for the bacteria towards the tissue surrounding the implant. Despite the close neighborhood of bacteria and myeloid cells, no phagocytosis had taken place. Only containment of bacteria to the pumps could be observed. However, activation of myeloid cells had taken place. Depletion of neutrophils led to an enhanced infection and to dissemination of the bacteria from the pumps to the neighboring tissue.
How do we get there?
Osmotic pumps will be exposed to bacterial cultures in order to pre-colonize the devices. The pumps will then be surgically administered subcutaneously to the mice and monitored for various time periods. A major technique to quantitate the bacteria in vivo will consist of non-invasive imaging of the bacteria. Thus, bacteria will be used that express the bacterial luciferase and can be detected within the living mouse using the IVIS systems. Plating under these circumstances will not provide valid data since we have noticed that part of the bacteria might be lost when the pumps are removed from the animals. On the other hand, non-invasive imaging allows following expansion or reduction of the number of bacteria on the device in real time and data can be acquired from the same individual mouse over time. The IVIS will also be employed to quantitate the bacteria under antibiotic therapy. The bacteria should be resilient against treatment since a mature biofilm should be protective. Modulated biofilms might be less protective. This would provide a simple read out system for our experiments. Analysis will also employ microscopic examination of the colonized pumps using light sheet a well as scanning electron microscopy. The latter will provide high resolution pictures while for light sheet microscopy biofilms can be stained using fluorescent antibodies against components of the biofilm or against the bacteria.
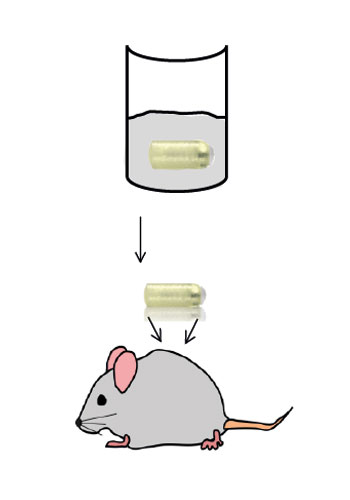
Mouse model for implant-associated infections.