Respiratory tract infections: What is the role of commensal bacteria in the respiratory tract?
What is this research project about?
The respiratory tract contains a well-documented bacterial community (microbiome) in the nasal cavity and nasopharynx, airways, and lung tissue. This lung microbiota successfully colonizes the respiratory tract, is tolerated by the host immune system, and can help combat invading external pathogens. Thus, the lung microbial ecosystem is maintained by the balance between transiently invading and selectively eliminated microorganisms. The bidirectional exchange between the lung microbiome and the host immune system plays a fundamental role in maintaining lung immune homeostasis. On the one hand, the inhabited lung microbiota can influence the maturation of the host immune system by producing numerous structural ligands and metabolites (such as lipopolysaccharide, peptidoglycan, short chain fatty acids (SCFA), secondary metabolites). On the other hand, the host innate and adaptive immune system can alter the lung microbiome by forming biophysical barriers, secreting cytokines, producing antimicrobial peptides, and recognizing resident and viable microbes. Therefore, many factors that cause dysbiosis of the lung microbiome can alter lung immune homeostasis and promote the occurrence and development of inflammation and respiratory disease.
What is the state of play?
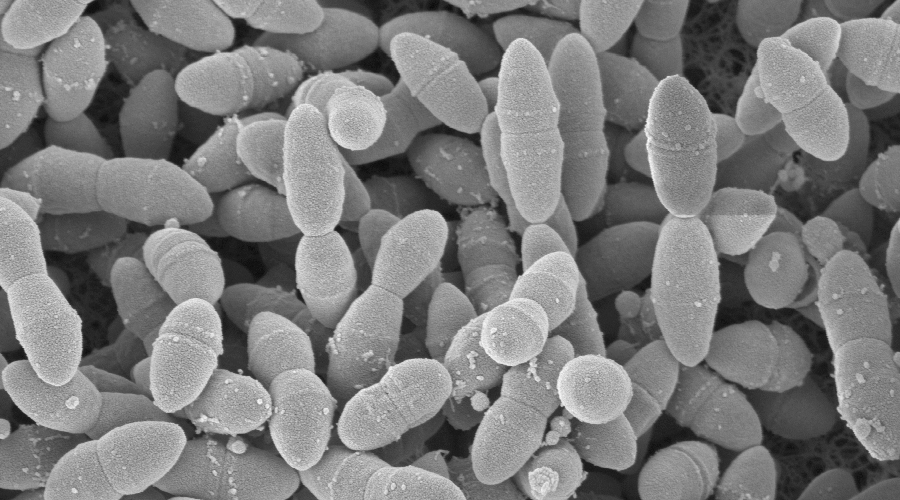
Several chronic inflammatory diseases of the lung have been found to be related to alterations in the composition of the airway microbiome. Moreover, the lung microbiota could be classified according to its predominance of proinflammatory bacteria such as Staphylococcus, Pseudomonas, and Haemophilus or of low-stimulatory bacteria such as Prevotella, Streptococcus, and Veillonella. However, the importance of the interplay between the lung microbiota and the airway epithelium is limited. Therefore, we hypothesized that the specific commensal microbiota of the lung may be important in protecting against infections with potentially pathogenic microbes in the airways.
In their previous RESIST project, Prof. Tümmler and Prof. Müller successfully analyzed the lung microbiome in various chronic lung diseases. They combined culture-dependent and -independent approaches to characterize the composition of the human lung microbiota, obtain representative cultured isolates and test their growth requirements, assess the temporal dynamics of these communities in the lung, and finally establish correlations with host health status.
How do we get there?
We will establish large-scale bacterial cultures based on samples from healthy individuals, covering a wide range of culture conditions. Subsequently, we will investigate whether commensals can also be cultured from the lungs of healthy individuals.
For further experiments, commensals that can be cultured and characterized by genome sequencing will be cultured either alone or with the pulmonary pathogens S. aureus, S. pneumonia, non-typable H. influenza, and M. catarrhalis in a dose- and time-dependent manner. We measure and compare the growth curves and metabolic activities. Each of the commensal bacteria will be co-cultured with each of the bacterial pathogens in lung epithelial cells and, where appropriate, we will subsequently perform further analyses to investigate the nature and extent of the impact of the bacterial interaction on the lung epithelium. For the commensal-induced interactions, we will further investigate whether specific chemical factors induced or produced by the bacteria could cause these changes.Our results provide an opportunity to identify potential factors that could be used to prevent or treat bacterial infections in the lung.