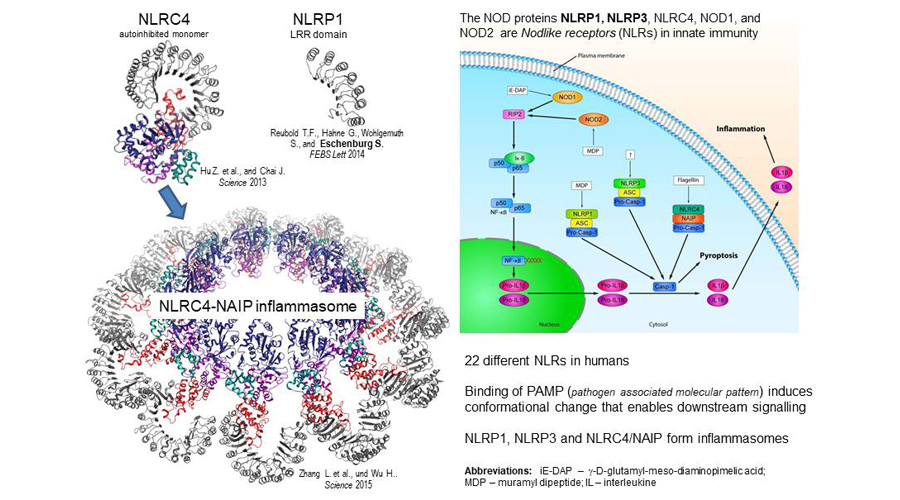
NLRs (nod-like receptors) initiate the defence against pathogens, but can also cause harm in humans. How can they be used for therapeutic purposes?
What is this research project about?
Nod-like receptors (NLRs) constitute a vital part of the innate immune system. These immune receptors recognize invading pathogens from within the cell and trigger an inflammatory reaction in the affected tissues via the activation and secretion of pro-inflammatory interleukines. NLRs are activated, amongst other conditions, upon infection with viruses and bacteria. Numerous viruses and bacteria in turn modulate the activity of NLRs to evade host defense or to support dissemination. Excessive activation of NLRs may severely aggravate the course of an infection by causing an overshooting inflammatory reaction and erroneous activation of NLRs is involved in numerous autoinflammatory disorders. Dampening of the activity of NLRs on the other hand underlies a substantial part of immunodeficiencies.
What’s the current status?
In recent years, NLRs and in particular the members of the NLRP subfamily of NLRs (NLRPs) have moved into the focus of medical research. Despite strong efforts to understand how NLRs work and how they may be targeted for therapeutic use, the knowledge about the molecular determinants of the activation and the function of NLRs is very limited. The same applies for a detailed knowledge about whether and how genetic variations in NLRs influence susceptibility to infections and the severity of the course of the infection. only a few substances are known that might act as inhibitors of individual NLRs.

How do we get there?
We combine a wide range of structural and cell biological methods in an integrative approach towards a fundamental understanding of NLRP function. To understand the dynamics of the activation of NLRPs, we structurally analyse the monomeric start states and the oligomeric end states of the activation process for several of the receptors. To depict the end states, the methods of choice are cryo-EM of the multimer in combination with X-ray crystallographic analysis of autoinhibited monomers or subdomains of NLRPs. The monomeric states of selected NLRPs, either in full-length or suitably truncated, will be analysed employing conventional crystallography or serial crystallography using a polychromatic synchrotron beam.
On the example of Kaposi sarcoma herpes virus (KSHV) and Herpes simplex virus 1 (HSV-1), we will elucidate how viruses modulate the activity of NLRPs. We recently confirmed in HEK293 cells that, upon infection with KSHV, the function of NLRP1 is blocked by direct interaction of the KSHV protein ORF63 with the receptor. We aim to determine the three-dimensional structures of KSHV-ORF63 and the related protein in HSV-1, in their isolated form as well as in complex with their target NLRPs. To identify SNPs in NLRPs that might influence the function of NLRPs we will use the information from the patient cohorts in RESIST.
For this purpose we will use data from the HSV and Zoster cohorts as well as data from the cohorts “Rheuma-VOR” and KFO 250. We will also use data from the RSV cohort IRIS and the cohort “Deutsches PID-NET”.
More information on the individual cohorts can be found here.