Why do infections with hanta- and arenaviruses progress so differently?
What is this research project about?
Infection with emerging zoonotic viruses such as members of the Arenavirus and Hantavirus families can result in significantly varying disease manifestations in humans, ranging from asymptomatic infections to highly pathogenic haemorrhagic fever. The determinants of disease are not well understood.
The aim of this project is to understand the host and viral factors underlying immune activation and variable disease manifestation and how these can be used for the development of novel and individualized antiviral treatment options.
What’s the current status?
While there are only up to approximately 100,000 recorded annual cases of hantavirus infections and up to 300,000 cases of Lassa fever, the most common disease associated with arenaviruses, the actual case numbers are most likely significantly higher due to a large number of asymptomatic or mild infections resembling other flu-like illnesses. For instance, it is estimated that 2–5% of the world population has at one point been infected with LCMV, a mostly apathogenic arenavirus.
Despite their potential prevalence and pathogenicity, there are currently no FDA- or EMA-approved antiviral treatments or vaccines available against highly pathogenic hantavirus or arenavirus infections, with only Ribavirin and Favipiravir having shown limited efficacy in some patients. Understanding the mechanisms of disease pathogenesis is essential for the development of novel and individualized antiviral treatment options.
How do we get there?
Our preliminary work shows that aberrant viral replication and the generation of defective viral genomes (DVGs) are a universal feature of negative-sense RNA viruses and that these DVGs are potent stimulants of the host interferon response. Thus, we would like analyze the mechanisms regulating DVG synthesis and the consequences thereof for viral pathogenesis of emerging RNA viruses.
Here, we are taking a multi-omics approach complemented with viral engineering and molecular biology tools to characterize the entire viral transcriptome/replicome and to identify the association between specific viral PAMPs and immune activation in vitro and in vivo.
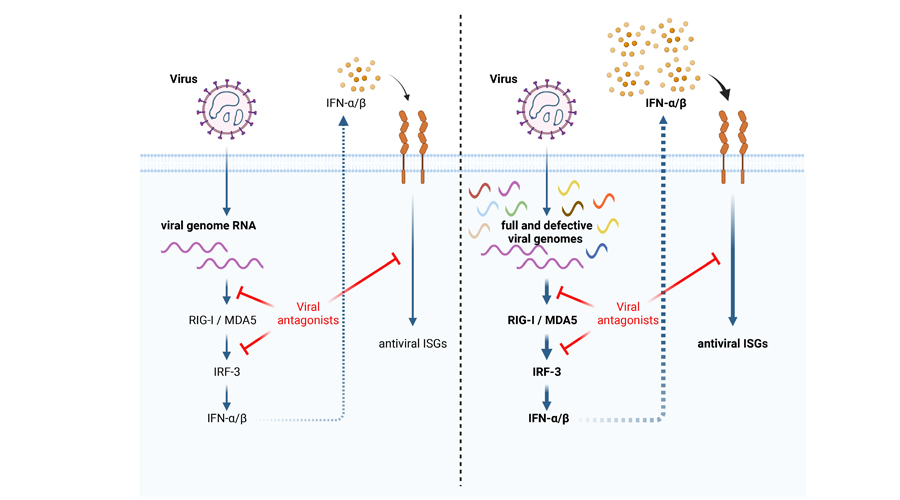
About the illustration: On the left, it can be seen that the RNA of the hantavirus (viral genome RNA) is sensed by cellular RNA sensors that trigger an interferon (IFN) response culminates in activation of antiviral genes. Viral IFN antagonists are able to inhibit IFN signalling. On the right, aberrant replication products / defective viral genomes accumulate to high numbers during viral replication. They are able to overwhelm the host’s innate immune system, leading to a prolonged and excessive IFN response. Thus, highly inflammatory and pathological infections can occur.