Publications 2025
A clinical protocol for a German birth cohort study of the Maturation of Immunity Against respiratory viral Infections (MIAI) C. R. Hartmann, R. Khan, J. Schöning, M. Richter, M. Willers, S. Pirr, et al. Front Immunol 2024 Vol. 15 Pages 1443665 Accession Number: 39355253 PMCID: PMC11442434 DOI: 10.3389/fimmu.2024.1443665
Oxidative phosphorylation is a key feature of neonatal monocyte immunometabolism promoting myeloid differentiation after birth G. Ehlers, A. M. Tödtmann, L. Holsten, M. Willers, J. Heckmann, J. Schöning, et al. Nat Commun 2025 Vol. 16 Issue 1 Pages 2239 Accession Number: 40050264 PMCID: PMC11885822 DOI: 10.1038/s41467-025-57357-w
The neonate respiratory microbiome S. Pirr, M. Willers and D. Viemann Acta Physiol (Oxf) 2025 Vol. 241 Issue 2 Pages e14266Accession Number: 39840649 PMCID: PMC11752418 DOI: 10.1111/apha.14266
Publications 2024
Postnatal supplementation with alarmins S100a8/a9 ameliorates malnutrition-induced neonate enteropathy in mice Lisa Perruzza # 1 2, Julia Heckmann # 3, Tanja Rezzonico Jost 4, Matteo Raneri 4, Simone Guglielmetti 5 6, Giorgio Gargari , Martina Palatella , Maike Willers , Beate Fehlhaber , Christopher Werlein , Thomas Vogl , Johannes Roth , Fabio Grassi , Dorothee Viemann Nat Commun. 2024 Oct 4;15(1):8623.
Early-life vitamin A treatment rescues neonatal infection-induced durably impaired tolerogenic properties of celiac lymph nodes. Mangge Zou, Joern Pezoldt, Juliane Mohr, Lars Philipsen, Andrea Leufgen, Vuk Cerovic, Carolin Wiechers, Marina Pils, Diego Ortiz, Lianxu Hao, Juhao Yang, Michael Beckstette, Aline Dupont, Mathias Hornef, Petra Dersch, Till Strowig, Andreas J. Müller, Jens Raila, Jochen Huehn, Cell Reports, Volume 43, Issue 5, 2024, 114153, ISSN 2211-1247.
The neonate respiratory microbiome S. Pirr, M. Willers and D. Viemann Acta Physiol (Oxf) 2025 Vol. 241 Issue 2 Pages e14266 Accession Number: 39840649 PMCID: PMC11752418 DOI: 10.1111/apha.14266
Publications 2023
Longitudinal development of the airway metagenome of preterm very low birth weight infants during the first two years of life. Rosenboom I, Pust MM, Pirr S, Bakker A, Willers M, Davenport CF, Wiehlmann L, Viemann D, Tümmler B. ISME Commun. 2023 Jul 20;3(1):75.
Silent neonatal influenza A virus infection primes systemic antimicrobial immunity. Heinemann AS, Stalp JL, Bonifacio JPP, Silva F, Willers M, Heckmann J, Fehlhaber B, Völlger L, Raafat D, Normann N, Klos A, Hansen G, Schmolke M, Viemann D. Front Immunol. 2023 Jan 24;14:1072142.
Insights into B cell ontogeny inferred from human immunology. Dirks J, Viemann D, Beyersdorf N, Härtel C, Morbach H. Eur J Immunol. 2023 Mar 11:e2250116.
Helicobacter spp. are prevalent in wild mice and protect from lethal Citrobacter rodentium infection in the absence of adaptive immunity. Bei Zhao, Lisa Osbelt, Till Robin Lesker, Marie Wende, Eric J.C. Galvez, Lisa Hönicke, Arne Bublitz, Marina C. Greweling-Pils, Guntram A. Grassl, Meina Neumann-Schaal, Till Strowig, Cell Reports, Volume 42, Issue 6, 2023
S100A9 is indispensable for survival of pneumococcal pneumonia in mice. Ostermann L, Seeliger B, David S, Flasche C, Maus R, Reinboth MS, Christmann M, Neumann K, Brand K, Seltmann S, Bühling F, Paton JC, Roth J, Vogl T, Viemann D, Welte T, Maus UA. PLoS Pathog. 2023 Jul 19;19(7):e1011493.
Publications 2022
Impact of gut microenvironment on epigenetic signatures of intestinal T helper cell subsets. Sasidharan Nair V, Heredia M, Samsom J, Huehn J. Immunol Lett. 2022 May 13;246:27-36.
Inflammatory perturbations in early life long-lastingly shape the transcriptome and TCR repertoire of the first wave of regulatory T cells Juhao Yang1,2†, Mangge Zou1†, Xiaojing Chu3, Stefan Floess1, Yang Li3, Michael Delacher4,5 and Jochen Huehn1,6* Front. Immunol., 31 August 2022
Postnatal expansion of mesenteric lymph node stromal cells towards reticular and CD34+ stromal cell subsets. Pezoldt J, Wiechers C, Zou M, Litovchenko M, Biocanin M, Beckstette M, Sitnik K, Palatella M, van Mierlo G, Chen W, Gardeux V, Floess S, Ebel M, Russeil J, Arampatzi P, Vafardanejad E, Saliba AE, Deplancke B, Huehn J. Nat Commun. 2022 Nov 24;13(1):7227.
The dark side of Tregs during aging. Palatella M, Guillaume SM, Linterman MA, Huehn J. Front Immunol. 2022 Aug 9
Publications 2021
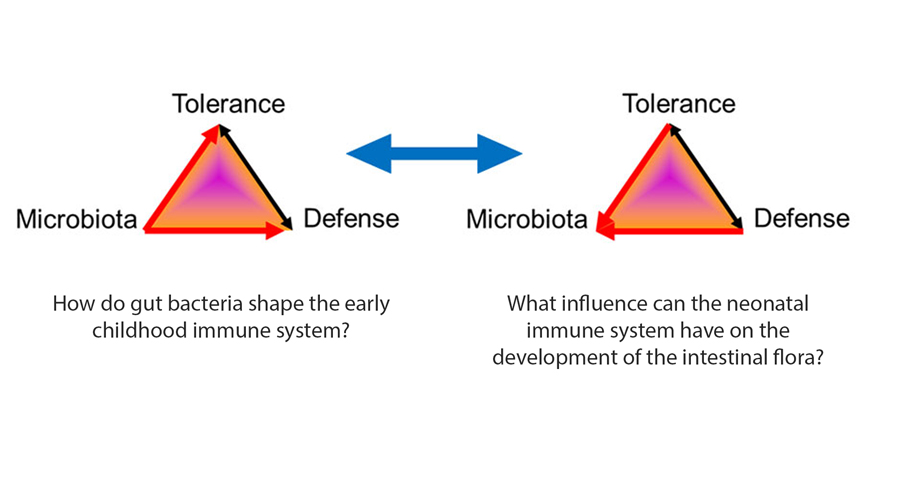
Role of the gut microbiota in airway immunity and host defense against respiratory infections. Willers M, Viemann D. Biol Chem. 2021 Oct 4. doi: 10.1515/hsz-2021-0281. Epub ahead of print. PMID: 34599869.
S100A8/A9 is the first predictive marker for neonatal sepsis. Pirr S, Dauter L, Vogl T, Ulas T, Bohnhorst B, Roth J, Viemann D.Clin Transl Med. 2021 Apr;11(4):e338. doi: 10.1002/ctm2.338.PMID: 33931974Free PMC article.No abstract available.
The microbiota is dispensable for the early stages of peripheral regulatory T cell induction within mesenteric lymph nodes. Wiechers C, Zou M, Galvez E, Beckstette M, Ebel M, Strowig T, Huehn J, Pezoldt J. Cell Mol Immunol. 2021 Mar 24. doi: 10.1038/s41423-021-00647-2. Online ahead of print. PMID: 33762684
Lymph node stromal cell subsets-Emerging specialists for tailored tissue-specific immune responses. Zou M, Wiechers C, Huehn J. Int J Med Microbiol. 2021 Apr;311(3):151492. doi: 10.1016/j.ijmm.2021.151492. Epub 2021 Feb 25. PMID: 33676241
Mesenteric Lymph Node Transplantation in Mice to Study Immune Responses of the Gastrointestinal Tract. Haroon Shaikh, Juan Gamboa Vargas, Zeinab Mokhtari, Katja J. Jarick, Maria Ulbrich, Josefina Peña Mosca, Estibaliz Arellano Viera, Caroline Graf, Duc-Dung Le, Katrin G. Heinze, Maike Büttner-Herold, Andreas Rosenwald, Joern Pezoldt, Jochen Huehn, Andreas Beilhack. Front Immunol. 2021; 12: 689896. Published online 2021 Jul 26.
Publications 2020
Evaluation of CD8 T cell killing models with computer simulations of 2-photon imaging experiments. Rastogi A, Robert PA, Halle S, Meyer-Hermann M.PLoS Comput Biol. 2020 Dec 28;16(12):e1008428. doi: 10.1371/journal.pcbi.1008428. Epub ahead of print. PMID: 33370254.
Influenza A viruses limit NLRP3-NEK7-complex formation and pyroptosis in human macrophages. Boal-Carvalho I, Mazel-Sanchez B, Silva F, Garnier L, Yildiz S, Bonifacio JP, Niu C, Williams N, Francois P, Schwerk N, Schöning J, Carlens J, Viemann D, Hugues S, Schmolke M. EMBO Rep. 2020 Nov 12;21(12):e50421. doi: 10.15252/embr.202050421. Epub ahead of print. PMID: 33180976; PMCID: PMC7726813.
Host Factors of Favorable Intestinal Microbial Colonization. Pirr S, Viemann D. Front Immunol. 2020 Oct 7;11:584288. doi: 10.3389/fimmu.2020.584288. PMID: 33117398; PMCID: PMC7576995.
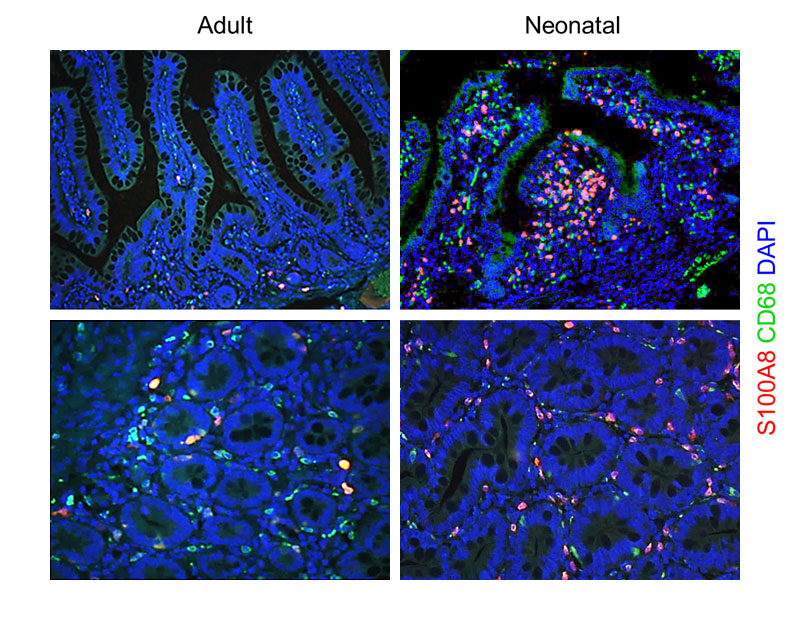
S100A8 and S100A9 Are Important for Postnatal Development of Gut Microbiota and Immune System in Mice and Infants. Willers M, Ulas T, Völlger L, Vogl T, Heinemann AS, Pirr S, Pagel J, Fehlhaber B, Halle O, Schöning J, Schreek S, Löber U, Essex M, Hombach P, Graspeuntner S, Basic M, Bleich A, Cloppenborg-Schmidt K, Künzel S, Jonigk D, Rupp J, Hansen G, Förster R, Baines JF, Härtel C, Schultze JL, Forslund SK, Roth J, Viemann D. Gastroenterology. 2020 Dec;159(6):2130-2145.e5. doi: 10.1053/j.gastro.2020.08.019. Epub 2020 Aug 15. PMID: 32805279.
Acute neonatal Listeria monocytogenes infection causes long-term, organ-specific changes in immune cell subset composition. Zou M, Yang J, Wiechers C, Huehn J. Eur J Microbiol Immunol (Bp). 2020 Jun 19;10(2):98–106. doi: 10.1556/1886.2020.00007. Epub ahead of print. PMID: 32644940; PMCID: PMC7391377.
S100-Alarmins Are Essential Pilots of Postnatal Innate Immune Adaptation. Viemann D. Front Immunol. 2020 Apr 30;11:688. doi: 10.3389/fimmu.2020.00688. PMID: 32425933; PMCID: PMC7203218.
IL-17 regulates DC migration to the peribronchial LNs and allergen presentation in experimental allergic asthma. Jirmo AC, Busse M, Happle C, Skuljec J, Dalüge K, Habener A, Grychtol R, DeLuca DS, Breiholz OD, Prinz I, Hansen G. Eur J Immunol. 2020 Jul;50(7):1019-1033. doi: 10.1002/eji.201948409. Epub 2020 Mar 29. PMID: 32142593.
Publications 2019
Microbiome Dependent Regulation of Tregs and Th17 Cells in Mucosa. Pandiyan P, Bhaskaran N, Zou M, Schneider E, Jayaraman S, Huehn J.
Front Immunol. 2019 Mar 8;10:426. eCollection 2019. Review.
Constitutive TNF-α signaling in neonates is essential for the development of tissue-resident leukocyte profiles at barrier sites. Bickes MS, Pirr S, Heinemann AS, Fehlhaber B, Halle S, Völlger L, Willers M, Richter M, Böhne C, Albrecht M, Langer M, Pfeifer S, Jonigk D, Vieten G, Ure B, von Kaisenberg C, Förster R, von Köckritz-Blickwede M, Hansen G, Viemann D. FASEB J. 2019 Oct;33(10):10633-10647. Epub 2019 Jun 29.
Injection of Antibodies against Immunodominant Epitopes Tunes Germinal Centers to Generate Broadly Neutralizing Antibodies. Meyer-Hermann M. Cell Rep. 2019 Oct 29;29(5):1066-1073.e5.
The role of Ames dwarfism and calorie restriction on gut microbiota. Wiesenborn DS, Gálvez EJC, Spinel L, Victoria B, Allen B, Schneider A, Gesing A, Al-Regaiey KA, Strowig T, Schäfer KH, Masternak MM. J Gerontol A Biol Sci Med Sci. 2019 Oct 30.
Impaired cellular energy metabolism in cord blood macrophages contributes to abortive response toward inflammatory threats. Dreschers S, Ohl K, Lehrke M, Möllmann J, Denecke B, Costa I, Vogl T, Viemann D, Roth J, Orlikowsky T, Tenbrock K. Nat Commun. 2019 Apr 11;10(1):1685. doi: 10.1038/s41467-019-09359-8. PMID: 30976008; PMCID: PMC6459909.