Which impact do intestinal bacteria and infections have on the fitness of immune cells?
What is this research project about?
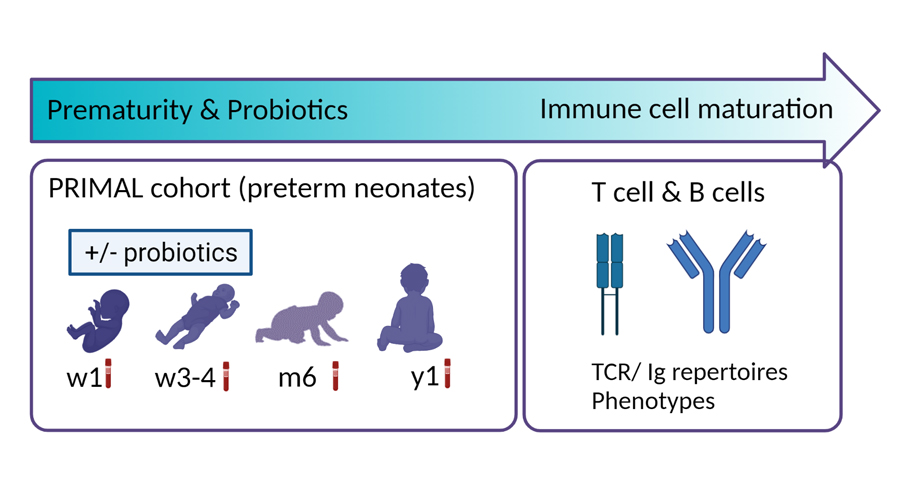
The project is based on the PRIMAL cohort. It is used to investigate the susceptibility to infections of premature babies in the first year of life after the administration of probiotics in the first weeks of life. We are investigating the maturation of T-cells and B-cells.
What is this research project about?
Infections are one of the main risks leading to neonatal morbidity and mortality of preterm infants. There is little information on how the immune cell maturation is impacted by health promoting factors like probiotica in preterm babies and how this differs from that of term infants.
In collaboration with Prof. Dr. Dorothee Viemann we aim at understanding immune cell maturation in a longitudinal cohort of preterm neonates that are supplemented with probiotics in the first weeks of life. In this project we focus on γδ T cells, αβ T cells and B cells, which are characterized by the expression of an antigen receptor. These immune cell subsets can either directly contribute to the immunity in neonates, but the majority undergoes a postnatal maturation driven by microbial challenges to become fully functional. This early age-related and environment-dependent imprinting on the antigen receptor repertoire and functionality of T cells and B cells might impact on the individual’s immune status and susceptibility towards infectious diseases during early life.
This project is based on the PRIMAL cohort. Prof. Viemann is heading and coordinating the PRIMAL cohort, a double-blinded study to understand the impact of probiotics supplements in the first weeks on disease susceptibility within the first year after preterm birth. We focus on understanding postnatal maturation of T cells and B cells.
What’s the current status?
Directly after birth, environmental factors such as the microbiota colonizing mucocutaneous barrier sites, but also occurring infections are important for the maturation of the children’s immune system. These host-pathogen interactions individually shape and imprint the immune system during early life. However, which environmental cues drive the adaptation of γδ T cells, αβ T cells and B cells with respect to their functionality and antigen-receptor repertoire after birth and how this might differ between preterm and term babies remains largely unclear.
To the picture: Age-dependent differences in antigen receptor repertoires. The Treemaps depict the abundance and distribution of individual T cell clones of γδ T cells in the peripheral blood of a newborn and an adult. Each square represents the abundance of a T cell clones according to the square size. We would like to decipher the impact of intrinsic factors, infections and microbiota on the postnatal maturation of antigen-receptor repertoires during early life.
How do we get there?
We have developed an mRNA-based high-throughput analysis technology and bioinformatics methods to analyze T-cell receptor repertoires within neonates and adults (Ravens et al., 2017). Initial data of defined patient cohorts indicates (i) a postnatal, extrathymic maturation of T cells (Ravens et al., 2017; DiLorenzo et al., 2019) and (ii) that viral infections leave a sustained footprint within the antigen receptor repertoire (Ravens et al., 2017; Ravens et al., 2018). The latter results propose that antigen receptor repertoires might serve as valuable biomarkers. More recently, we found that defined T cells develop in the early human thymus and immediately expand in response to the microbiota after preterm birth (Ravens, Fichtner et al., 2020, Tan, Fichtner et al., 2021).
Altogether, these findings reflect the developmental plasticity and suggest options for an iatrogenic imprinting of innate and adaptive immunity to promote immune maturation and adaptation during early life in individuals at high risk for infection- and immune-mediated diseases. Together with RG Viemann (RESIST project B1) we work on how defined factors influence the functional maturation of T cells and B cells within preterm infants and how the expressed antigen-receptor repertoire is shaped during early life.