What influence does genetic diversity have on the bacterial phenotype?
What is this research project about?
We usually think of genetic differences in the context of human genetics. And so we are familiar with the fact that two people differ on average at about five million sites in the genome – that is, at only about 0.8 per cent of the entire genome. But genetic variation in bacteria has a completely different dimension: For example, two strains of Escherichia coli can differ in up to 60 percent of their genetic content.
This means that each bacterial species has not only a single genome, but also a diverse ensemble of gene combinations called a pangenome. We want to investigate the influence of this great genetic diversity on bacterial phenotypes.
What’s the current status?
Understanding how variations in genotype lead to variations in phenotype is an old topic in molecular biology. We are exploring how to better predict the phenotypic consequences of genetic variants in bacteria. This can be done with a combination of “forward-looking” mechanistic models, and “backward-looking” statistical genomics models.
Certain bacterial species, such as E.coli, have so-called open pangenomes. This means that new genes (“accessory genes”) are discovered when the genomes of new isolates are sequenced. Current functional genomics techniques cannot scale up fast enough to keep up with the flood of genes with unknown function that are being discovered every day.
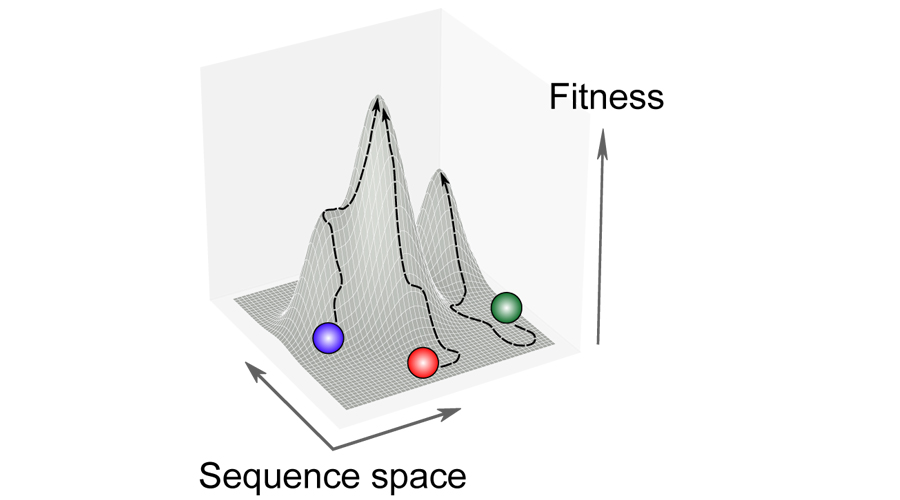
Studying how different bacterial strains adapt to the same selective pressure (e.g. antimicrobials).
How do we get there?
It has been shown that the genetic variability between bacterial strains also affects genes that are indispensable for growth and reproduction. We suspect that these differences could also influence the ability to adapt to selection pressures, such as the use of antimicrobials. As we explore the influence of bacterial pangenomes on clinically relevant phenotypes, we are also investigating whether different genetic backgrounds make people more or less susceptible to developing antibiotic resistance.
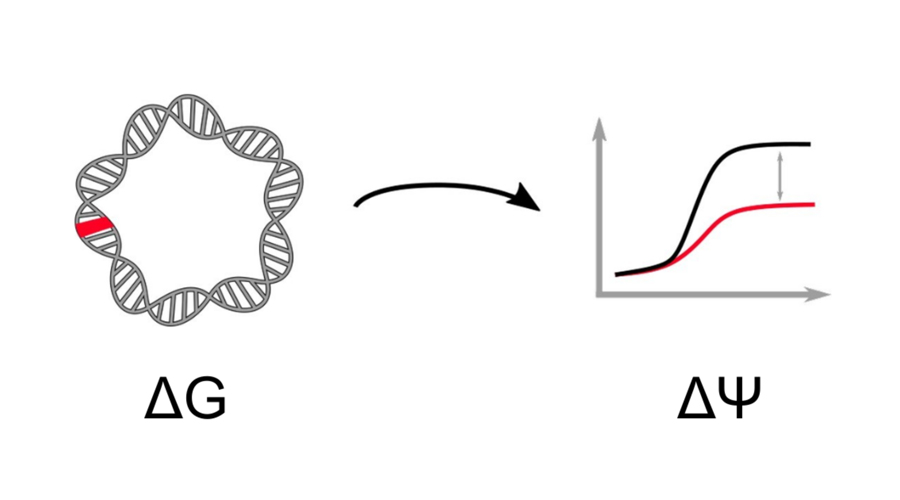
Computational and experimental approaches to link genotypic to phenotypic variation.