How can a precise characterisation of microbial communities result in the development of new strategies against biofilm-associated infections?
What is this research project about?
Bacteria in biofilms are embedded in a self-produced extracellular matrix and exhibit an increased resistance to adverse conditions. In the human host, biofilm bacteria are responsible for persistent infections and efficiently withstand antibiotic treatment and the host immune response. Once a bacterial biofilm infection is established it becomes very difficult to eradicate, even in the absence of genotypic resistance. Biofilm infections affect millions of people and every year chronic infections in patients due to biofilm formation are a multi-billion Euro burden to national healthcare systems. With progress of medical sciences, more and more indwelling devices for the purpose of medical treatments and foreign body implants are applied. Infection continues to be a major complication of their use. Also, there are biofilm infections not associated with foreign bodies, such as chronic infections of the lungs of cystic fibrosis patients and of patients with chronic obstructive pulmonary diseases. Ongoing inflammation and changes in the structure and function of the affected tissue largely determine morbidity and mortality in these patients.
We want to identify biomarkers whose presence is correlated with the resistance of the Pseudomonas aeruginosa biofilm and genetic / metabolic patterns and which characterize the switch to the establishment of pathogenic biofilms on implants. This will serve to develop a diagnostic tool for biofilm resistance and disease excitation Profiling and innovative treatment strategies targeting biofilm resistance mechanisms.
What’s the current status?
Although chronic biofilm-associated infections have been extensively studied, there are many open questions and the general recalcitrance of biofilm-grown bacteria is only incompletely understood. The successful use of antibiotics to eradicate biofilm-associated infection relies on our ability to overcome several main problems. First, for a more targeted anti-biofilm therapy, knowledge on the biofilm-specific resistance profile of individual bacterial isolates is essential, as well as knowledge on when natural colonizing bacterial communities transition to pathogenic biofilms e.g. on implants. In addition, new therapy options will have to be developed to overcome the second limitation of current treatment, which is the general recalcitrance of biofilm populations.
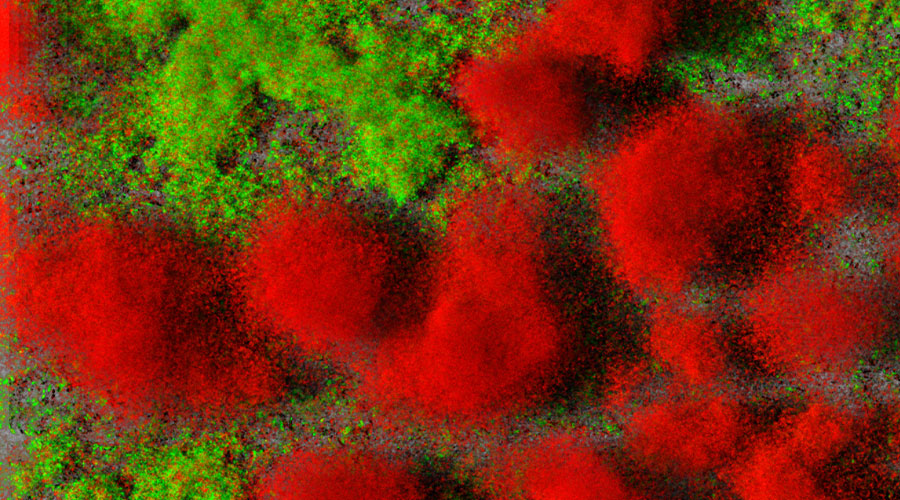
Biofilms of clinical idolates of Pseudomonas aeruginosa, living cells are green, dead cells are red. Source: TWINCORE / Jann Thöming
How do we get there?
Our research groups have established extensive expertise in the analysis of the structure, assembly and microbiological diversity of medical biofilms, and have applied methodologies including DNA/RNA sequencing and machine learning approaches to describe the genomic and transcriptional landscape of infecting pathogens in vitro and ex vivo. Within RESIST we want to transfer gained knowledge and experience from our work on bacterial biofilms and establish a genome-based prediction of bacterial phenotypes by integrating complex OMICS-data also using machine learning classifiers, phylogenomic clustering and feature selection techniques.

Biofilms of clinical idolates of Pseudomonas aeruginosa, living cells are green, dead cells are red. Source: TWINCORE / Jann Thöming